drought |
effect of drought on histone modification in plants |
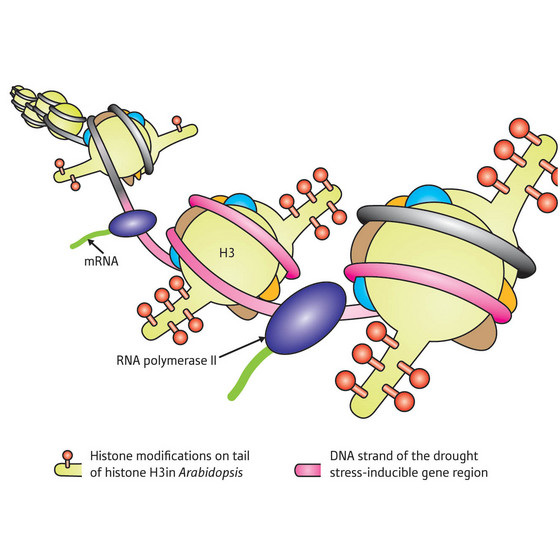
Drought stress affects plant growth and survival. Gene regulatory networks associated with the drought stress response in plants have been studied by analyzing drought stress responsive genes that encode functional and regulatory proteins, such as transcription factors.The expression of drought stress-responsive genes is positively correlated with the intensity of drought stress. Under strong drought conditions, the histone modifications H3K4me3 and H3K9ac on drought stress-upregulated genes, such as RD20 and RD29A, were more highly enriched than under moderate drought conditions. Furthermore, under moderate drought conditions, there was little nucleosome loss from the RD29A region, while under strong drought conditions, notable nucleosome loss occurred in the same gene region. These results indicated that epigenetic responsiveness depended on the intensity of the drought stress[K. Sano et. al. 2015]. More recently, it was
found that histone modifications on histone H3 were altered
with gene activation on the coding regions of drought
stress-responsive genes under drought stress conditions
Histone methylation and DNA methylation happens together. In addition to H3K9 methylation and histone methyltransferases,other histone modifications and chromatin modifiers can influence the maintenance of DNA methylation.[T. Kanno et. al. 2012]
Histone acetylation status has also been correlated with drought stress and ABA responses in plants. For example, in response to drought stress, the histone acetylation level increased on the drought-responsive genes, such as RD20, RD29A, and RD29B and H3K9ac was enriched rapidly in these gene regions. Interestingly, specific patterns of histone acetylation were found on each drought-responsive gene, suggesting that differences in histone modification may contribute to the responsiveness of these genesThe HD2-type HDACs, HD2A (HDT1), HD2B (HDT2), HD2C (HDT3), and HD2D (HDT4) belong to a plant-specific HDAC family in Arabidopsis, rice, barley, and tomato AtHD2C-overexpressing Arabidopsis plants showed ABA insensitivity, reduced transpiration, and enhanced tolerance to drought and salt stresses. In barley, the expression of HD2 genes responded to stress-related plant hormones such as ABA, jasmonic acid (JA), and salicylic acid (SA) . These results suggested that HD2 genes play roles in resistance to abiotic and biotic stresses in monocot and dicot plants.[K. Sano et. al 2015]. The reversible acetylation and deacetylation of specific lysine residues on core histones are catalysed by histone acetyltransferases and histone deacetylases (HDAs). HDA6 is an RPD3-type histone deacetylase in Arabidopsis. The Arabidopsis HDA6 mutant, axe1-5, and HDA6 RNA-interfering plants displayed a phenotype that was hypersensitive to ABA and drought stress[Keqiang Wu et. al. 2010].
There are two types of regulatory mechanisms of nucleosome occupancy in response to drought stress. The first type is that histone loss occurs on the promoter and coding regions in response to drought stress. RD20 and RAP2.4 genes fit this case. Generally, nucleosome occupancy on the coding regions is inversely correlated with RNAPII association and transcriptional activity. The second type is that nucleosome density is low on the promoter regions and histone loss does not occur on the coding regions in response to drought stress. [Jong-Myong Kim et. al. 2008]
A direct connection between the temperature-sensing H2A.Z at transcription-competent start sites of genes and the heat-induced loss of nucleosomes from heterochromatic repeats is unlikely as DNA methylation typical for the latter is mutually exclusive with H2A.Z domains. However, both responses have in common that the removal of histones does not increase expression of all genes equally and therefore is not
sufficient for transcriptional activation. The occurrence of multiple histone variants, modifications, chaperones and different nucleosome loading make it likely that chromatin dynamics upon stress are the result of a complex interplay between physical factors, their perception, pre-existing chromatin structure and maintenance mechanisms[Ales and Ortrum 2013].
By these way histone variants modify due to drought stress.
found that histone modifications on histone H3 were altered
with gene activation on the coding regions of drought
stress-responsive genes under drought stress conditions
Histone methylation and DNA methylation happens together. In addition to H3K9 methylation and histone methyltransferases,other histone modifications and chromatin modifiers can influence the maintenance of DNA methylation.[T. Kanno et. al. 2012]
Histone acetylation status has also been correlated with drought stress and ABA responses in plants. For example, in response to drought stress, the histone acetylation level increased on the drought-responsive genes, such as RD20, RD29A, and RD29B and H3K9ac was enriched rapidly in these gene regions. Interestingly, specific patterns of histone acetylation were found on each drought-responsive gene, suggesting that differences in histone modification may contribute to the responsiveness of these genesThe HD2-type HDACs, HD2A (HDT1), HD2B (HDT2), HD2C (HDT3), and HD2D (HDT4) belong to a plant-specific HDAC family in Arabidopsis, rice, barley, and tomato AtHD2C-overexpressing Arabidopsis plants showed ABA insensitivity, reduced transpiration, and enhanced tolerance to drought and salt stresses. In barley, the expression of HD2 genes responded to stress-related plant hormones such as ABA, jasmonic acid (JA), and salicylic acid (SA) . These results suggested that HD2 genes play roles in resistance to abiotic and biotic stresses in monocot and dicot plants.[K. Sano et. al 2015]. The reversible acetylation and deacetylation of specific lysine residues on core histones are catalysed by histone acetyltransferases and histone deacetylases (HDAs). HDA6 is an RPD3-type histone deacetylase in Arabidopsis. The Arabidopsis HDA6 mutant, axe1-5, and HDA6 RNA-interfering plants displayed a phenotype that was hypersensitive to ABA and drought stress[Keqiang Wu et. al. 2010].
There are two types of regulatory mechanisms of nucleosome occupancy in response to drought stress. The first type is that histone loss occurs on the promoter and coding regions in response to drought stress. RD20 and RAP2.4 genes fit this case. Generally, nucleosome occupancy on the coding regions is inversely correlated with RNAPII association and transcriptional activity. The second type is that nucleosome density is low on the promoter regions and histone loss does not occur on the coding regions in response to drought stress. [Jong-Myong Kim et. al. 2008]
A direct connection between the temperature-sensing H2A.Z at transcription-competent start sites of genes and the heat-induced loss of nucleosomes from heterochromatic repeats is unlikely as DNA methylation typical for the latter is mutually exclusive with H2A.Z domains. However, both responses have in common that the removal of histones does not increase expression of all genes equally and therefore is not
sufficient for transcriptional activation. The occurrence of multiple histone variants, modifications, chaperones and different nucleosome loading make it likely that chromatin dynamics upon stress are the result of a complex interplay between physical factors, their perception, pre-existing chromatin structure and maintenance mechanisms[Ales and Ortrum 2013].
By these way histone variants modify due to drought stress.
Disclaimer: This website is a part of academics and used only for educational purpose.
